R code for example in Chapter 7:
Analyzing proportions
New methods on this page
These are the core commands that produce the new methods described in Chapter 7. The variable names are plucked from the examples further below.
- Calculating binomial probabilities
dbinom(6, size = 27, prob = 0.25)- Binomial test
binom.test(10, n = 25, p = 0.061)- Install an R package
install.packages("binom", dependencies = TRUE)- Agresti-Coull 95% confidence interval for the proportion
binom.confint(30, n = 87, method = "ac")Other new methods
Sampling distribution of a proportion by repeated sampling from a known population.
Examples
Table 7.1-1 and Figure 7.1-1. Binomial distribution with n = 27 and p = 0.25
Table and histogram of binomial probabilities. Uses the data from Chapter 6 on the genetics of mirror-image flowers.
Calculate a binomial probability, the probability of obtaining X successes in n trials when trials are independent and probability of success p is the same for every trial. The probability of getting exactly 6 left-handed flowers when n = 27 and p = 0.25 is
dbinom(6, size = 27, prob = 0.25)## [1] 0.171883Table of probabilities for all possible values for the number of left-handed flowers out of 27.
xsuccesses <- 0:27
probx <- dbinom(xsuccesses, size = 27, prob = 0.25)
probTable <- data.frame(xsuccesses, probx)
probTable## xsuccesses probx
## 1 0 4.233057e-04
## 2 1 3.809751e-03
## 3 2 1.650892e-02
## 4 3 4.585812e-02
## 5 4 9.171623e-02
## 6 5 1.406316e-01
## 7 6 1.718830e-01
## 8 7 1.718830e-01
## 9 8 1.432358e-01
## 10 9 1.007956e-01
## 11 10 6.047736e-02
## 12 11 3.115500e-02
## 13 12 1.384667e-02
## 14 13 5.325641e-03
## 15 14 1.775214e-03
## 16 15 5.128395e-04
## 17 16 1.282099e-04
## 18 17 2.765311e-05
## 19 18 5.120947e-06
## 20 19 8.085705e-07
## 21 20 1.078094e-07
## 22 21 1.197882e-08
## 23 22 1.088984e-09
## 24 23 7.891188e-11
## 25 24 4.383993e-12
## 26 25 1.753597e-13
## 27 26 4.496403e-15
## 28 27 5.551115e-17Histogram of binomial probabilities for the number of left-handed flowers out of 27. This illustrates the full binomial distribution when n = 27 and p = 0.25.
barplot(height = probx, names.arg = xsuccesses, space = 0, las = 1,
ylab = "Probability", xlab = "Number of left-handed flowers")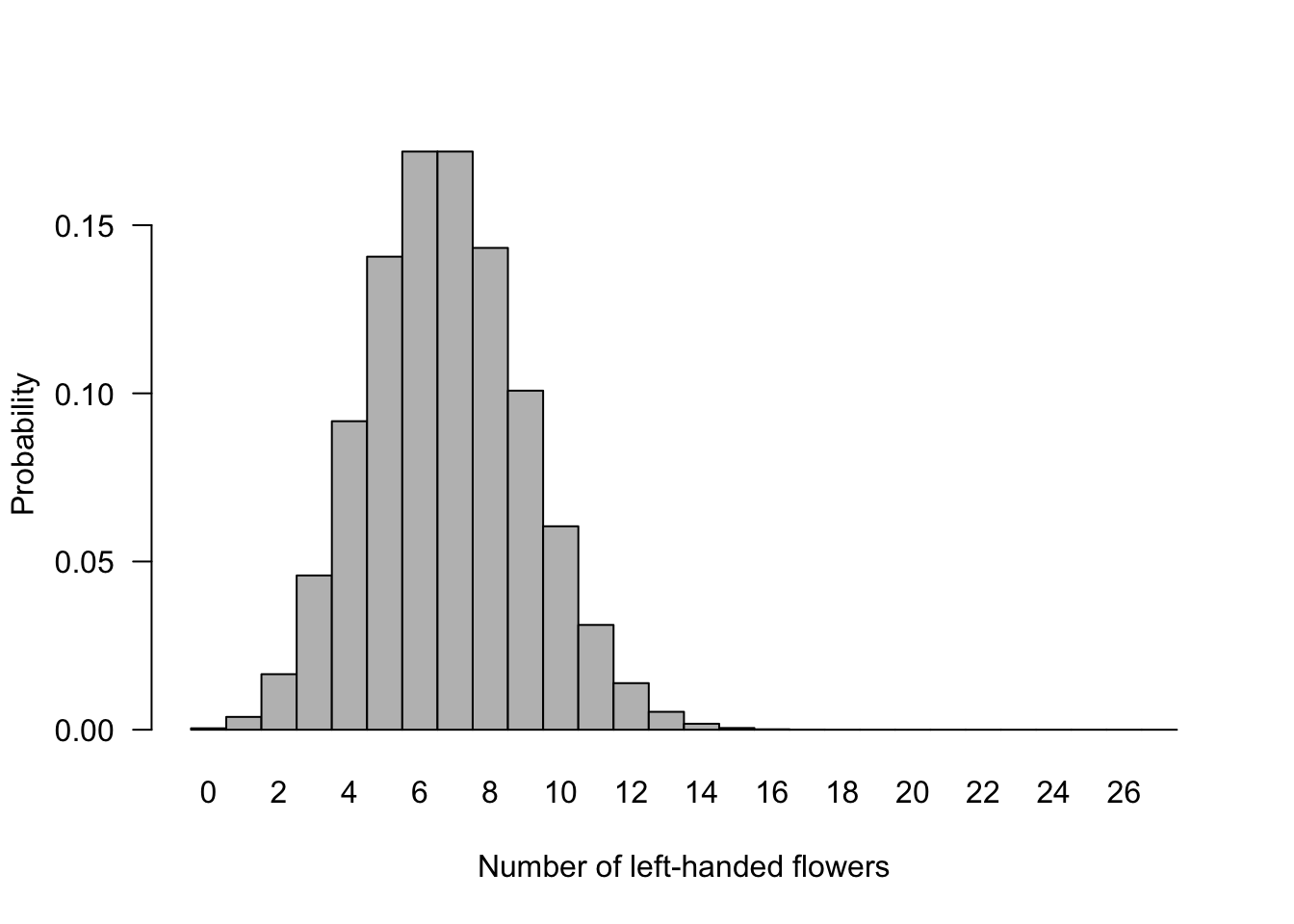
Figure 7.1-2. Sampling distribution of a binomial proportion
Compare sampling distributions for the proportion based on n = 10 and n = 100.
Take a large number of random samples of n = 10 from a population having probability of success p = 0.25. Convert to proportions by dividing by the sample size. Do the same for the larger sample size n = 100. The following commands use 10,000 random samples.
successes10 <- rbinom(10000, size = 10, prob = 0.25)
proportion10 <- successes10 / 10
successes100 <- rbinom(10000, size = 100, prob = 0.25)
proportion100 <- successes100 / 100Plot and visually compare the sampling distributions of the proportions based on n = 10 and n = 100. The par(mfrow = c(2,1)) command sets up a graph window that will plot both graphs arranges in 2 rows and 1 column.
par(mfrow = c(2,1))
hist(proportion10, breaks = 10, right = FALSE, xlim = c(0,1),
xlab = "Sample proportion")
hist(proportion100, breaks = 20, right = FALSE, xlim = c(0,1),
xlab = "Sample proportion")
par(mfrow = c(1,1))Commands for a fancier plot are here.
oldpar <- par(no.readonly = TRUE) # make backup of default graph settings
par(mfrow = c(2,1), oma = c(4, 0, 0, 0), mar = c(1, 6, 4, 1)) # adjust margins
saveHist10 <- hist(proportion10, breaks = 10, right = FALSE, plot = FALSE)
saveHist10$counts <- saveHist10$counts/sum(saveHist10$counts)
plot(saveHist10, col = "firebrick", las = 1, cex.lab = 1.2,
ylim = c(0,0.3), xlim = c(0,1), ylab = "Relative frequency",
xlab = "", main = "")
text(x = 1, y = 0.25, labels = "n = 10", adj = 1, cex = 1.1)
saveHist100 <- hist(proportion100, breaks = 40, right = FALSE, plot = FALSE)
saveHist100$counts <- saveHist100$counts/sum(saveHist100$counts)
plot(saveHist100, col = "firebrick", las = 1, cex.lab = 1.2,
ylim = c(0,0.1), xlim = c(0,1), ylab = "Relative frequency",
xlab = "", main = "")
text(x = 1, y = 0.08, labels = "n = 100", adj = 1, cex = 1.1)
mtext("Proportion of successes", side = 1, outer = TRUE, padj = 2)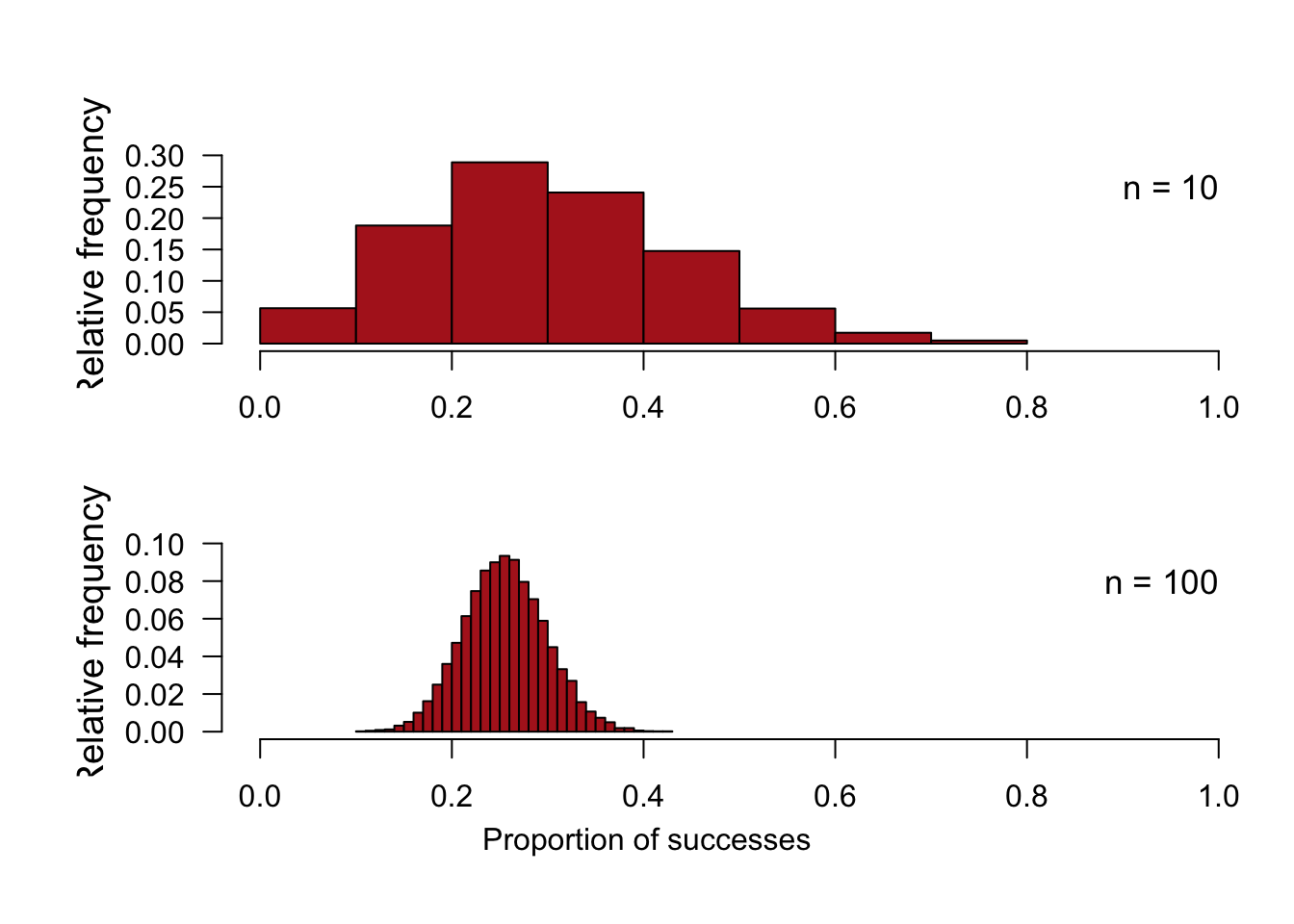
par(oldpar) # Revert to backup graph settingsExample 7.3. Sex and the X chromosome
The binomial test, used to test whether spermatogenesis genes in the mouse genome occur with unusual frequency on the X chromosome.
Read and inspect the data. Each row in the data file represents a different spermatogenesis gene.
mouseGenes <- read.csv("../Data/chapter07/chap07e2SexAndX.csv")
head(mouseGenes)## chromosome onX
## 1 4 no
## 2 4 no
## 3 6 no
## 4 6 no
## 5 6 no
## 6 7 noTabulate the number of spermatogenesis genes on the X-chromosome and the number not on the X-chromosome.
table(mouseGenes$onX)##
## no yes
## 15 10Calculate the binomial probabilities of all possible outcomes under the null hypothesis (Table 7.2-1). Under the binomial distribution with n = 25 and p = 0.061, the number of successes can be any integer between 0 and 25.
xsuccesses <- 0:25
probx <- dbinom(xsuccesses, size = 25, prob = 0.061)
data.frame(xsuccesses, probx)## xsuccesses probx
## 1 0 2.073193e-01
## 2 1 3.367007e-01
## 3 2 2.624760e-01
## 4 3 1.307255e-01
## 5 4 4.670757e-02
## 6 5 1.274386e-02
## 7 6 2.759585e-03
## 8 7 4.865905e-04
## 9 8 7.112305e-05
## 10 9 8.727323e-06
## 11 10 9.071211e-07
## 12 11 8.035781e-08
## 13 12 6.090306e-09
## 14 13 3.956429e-10
## 15 14 2.203032e-11
## 16 15 1.049510e-12
## 17 16 4.261188e-14
## 18 17 1.465509e-15
## 19 18 4.231266e-17
## 20 19 1.012696e-18
## 21 20 1.973624e-20
## 22 21 3.052667e-22
## 23 22 3.605629e-24
## 24 23 3.055193e-26
## 25 24 1.653947e-28
## 26 25 4.297797e-31Use these probabilities to calculate the P-value corresponding to an observed 10 spermatogenesis genes on the X chromosome. Remember to multiply the probability of 10 or more successes by 2 for the two-tailed test result.
2 * sum(probx[xsuccesses >= 10])## [1] 1.987976e-06For a faster result, try R’s built-in binomial test. The resulting P-value is slightly different from our calculation. In the book, we get the two-tailed probability by multiplying the one-tailed probability by 2. As we say on page 188, computer programs may calculate the probability of extreme results at the “other” tail with a different method. The output of binom.test includes a confidence interval for the proportion using the Clopper-Pearson method, which is more conservative than the Agresti-Coull method.
binom.test(10, n = 25, p = 0.061)##
## Exact binomial test
##
## data: 10 and 25
## number of successes = 10, number of trials = 25, p-value =
## 9.94e-07
## alternative hypothesis: true probability of success is not equal to 0.061
## 95 percent confidence interval:
## 0.2112548 0.6133465
## sample estimates:
## probability of success
## 0.4Example 7.2. Radiologists’ missing sons
Standard error and 95% confidence interval for a proportion using the Agresti-Coull method for the confidence interval.
Read and inspect the data.
radiologistKids <- read.csv("../Data/chapter07/chap07e3RadiologistOffspringSex.csv")
head(radiologistKids)## offspringSex
## 1 male
## 2 male
## 3 male
## 4 male
## 5 male
## 6 maleFrequency table of female and male offspring number.
table(radiologistKids$offspringSex)##
## female male
## 57 30Calculate the estimated proportion of offspring that are male, and the total number of radiologists.
n <- sum(table(radiologistKids$offspringSex))
n## [1] 87pHat <- 30 / n
pHat## [1] 0.3448276Standard error of the sample proportion.
sqrt( (pHat * (1 - pHat))/n )## [1] 0.0509588Agresti-Coull 95% confidence interval for the population proportion.
pPrime <- (30 + 2)/(n + 4)
pPrime## [1] 0.3516484lower <- pPrime - 1.96 * sqrt( (pPrime * (1 - pPrime))/(n + 4) )
upper <- pPrime + 1.96 * sqrt( (pPrime * (1 - pPrime))/(n + 4) )
c(lower = lower, upper = upper)## lower upper
## 0.2535425 0.4497542Agresti-Coull 95% confidence interval for the population proportion using the binom package. To use this package you will need to install it (this needs to be done only once per computer) and load it using the library command (this needs to be done once per R session). The confidence interval from the binom package will be very slightly different from the one you calculated above because the formula we use takes a slight shortcut.
install.packages(“binom”, dependencies = TRUE)
library(binom)
binom.confint(30, n = 87, method = "ac")## method x n mean lower upper
## 1 agresti-coull 30 87 0.3448276 0.2532164 0.4495625